Specification of Cell Types¶
Individual cells are the elementary unit of our models. Nevertheless, they are not represented as homogeneous bags in our framework. Instead we use Stochastic P-systems (SP-systems) [Romero-Campero2009] as the computational abstraction for an individual cell. The specification of an individual cell consists of the following three main components:
- The molecular species (genes, RNAs, proteins, signals etc.) present in the specific cell type are represented using string-objects like proteinGFP, signal3OC6, etc.
- The compartments (cytoplasm, nucleus, mitochondria, etc) of individual cells as well as some relevant regions related to individual cells (cell surface, media surronding a cell, etc.) are defined using membranes. The inclusion of compartments inside other compartements as in the case of the nucleus being inside the cytoplasm can also be specified.
- The characteristic molecular interactions taking place inside or between specific compartments are described using rules where the reactant and product molecules are specified as well as the compartment involved in the interaction and a stochastic constant used to compute the probability of applying each rule and the time elapsed between rule applications.
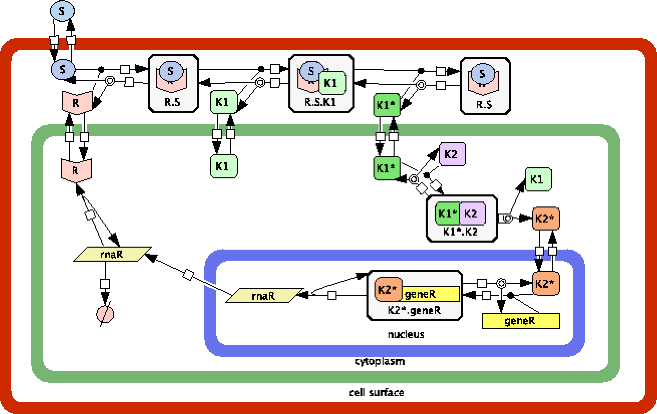
Example of an SP-system model of a cell type with three compartments, cell surface, cytoplasm and nucleus, string-objects representing molecular species as receptors, signals and genes and molecular interactions specified as rules describing a signal transduction pathway.
The specification of the components of a model of an individual cell using the Infobiotics modelling language is described in the following sections: