Model Specification¶
Infobiotics Workbench features the Infobiotics Language (IBL) - a the domain-specific language for synthetic biology, which allow the integration of various computational aspects into a single specification file. IBL defines a simple combined grammar for modelling, verification and biocompilation statements, facilitating an iterative workflow on building a synthetic biological device by using directives on what simulations to run, what verifications to compute and what synthetic circuits to biocompile.
Overview¶
Infobiotics Language allow biological entities to be designed using robut encapsulation and hierarchical structure mechanisms, featuring the following concepts:
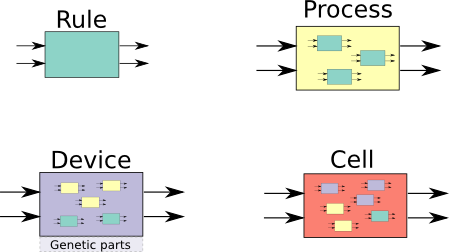
- Molecular species - any biological entity, from PROTEIN to DNA to MOLECULE;
- RULE - used to define a transformation, most often a chemical transformation;
- PROCESS - collection of RULEs. They make it easier to re-use rules and typically represent a more abstract process, e.g. constitutive protein expression;
- DEVICE - collection of PROCESSes and RULEs. They refer to device in the synthetic biology sense: a piece of DNA made up of parts;
- SYSTEM - contains DEVICEs and RULEs;
- PLASMID - collection of SYSTEMs, DEVICEs and RULEs. They are useful when more than one type of plasmid is used in the cell;
- CHROMOSOME - collection of SYSTEMs, DEVICEs and RULEs;
- CELL - biological cells. They are a collection of PLASMIDs, SYSTEMs, DEVICEs, PROCESSes and RULEs;
- REGION - a location in the lattice, defining the geometry of the simulation. They contain molecular species, CELLs, PROCESSes and RULEs.
Biological Parts¶
IBL allows the declaration of biological parts:
- genetic molecular components, e.g. PROMOTER, GENE, RIBOSOME, TERMINATOR, CLONING SITE, DNA, RNA and RBS, with optional declaration of sequences either in online repositories or in a built-in database;
- non-genetic molecular components, e.g. PROTEIN, MOLECULE.
Infobiotics Language supports the specification of concentrations in SI units and automatically converts between units, where possible. Concentrations can be specified in molar (i.e. moles/liter), millimolar, micromolar, and nanomolar units. Alternatively, concentration can be also be specified as an amount of molecules, which can be interconverted with molar concentration using a defined cell volume (which defaults to one femtoliter).
The code snippet below illustrates the syntax used for declaring various biological parts:
1 2 3 | promoter = PROMOTER ("URI=http://biobricks.org/...")
gene = GENE ("URI=http://parts.igem.org/...")
LacI = PROTEIN (concentration=10 uM)
|
Thus, declared biological parts can be connected to each other via reaction rules, which can be declared as either reversible or irreversible. Since complexification is particularly important in biological systems, e.g. in the quaternary structure of proteins or genetic regulation, complexes can be used in rules without the need to explicitly declare them.
PROCESSes¶
Being designed for a scalability, modularity and reusability, IBL allows for the declaration of PROCESS entities, which group together a set of rules involving abstract biological entities. After declaration, PROCESSes can be instantiated in specific components. For example, one can specify a process that collects all of the reactions that constitute general regulated gene expression, which then can be instantiated for different transcription factors and genes. In this manner, processes introduce a layer of modularity and abstraction similar to function declarations in imperative programming languages.
Infobiotics Language supports the specification of reaction rate constants in SI units, e.g. per second, per minute and per hour in case of unimolecular reactions, or per mole per second etc. for bimolecular reactions.
The example below illustrates the syntax used for declaring a process, containing:
- The declaration of LacI - protein with a concentration of 10 micromoles per liter;
- The declaration of Plac - promoter, one molecule (by default);
- A reversible rule for the dimerization of two LacI units, specifying also the forward and backward rate constants;
- A reversible rule for the binding of the LacI dimer to the Plac promoter, together with its corresponding forward and backward rate constants.
1 2 3 4 5 6 7 8 9 10 11 12 13 | define ConstitutiveProteinExpression typeof PROCESS {
LacI = PROTEIN (concentration=10 uM)
Plac = PROMOTER()
RULE lacI_dimerization: LacI + LacI <-> LacI~LacI
lacI_dimerization.forwardRate = 1e0 M-1 min-1
lacI_dimerization.backwardRate = 125 min-1
RULE lac_repression: Plac + LacI~LacI <-> Plac~LacI~LacI
lac_repression.forwardRate = 1e0 min-1
lac_repression.backwardRate = 10 min-1
}
|
DEVICEs¶
Infobiotics Language provides the means to group molecules, rules, and process instantiations into DEVICE entities, allowing the designer to capture the action of an entire genetic device, e.g. a strand of DNA containing several promoters, genes, and their encoded proteins.
The code snippet below shows the syntax required for declaring a device:
1 2 3 4 5 6 7 8 | operon = DEVICE (
parts =[PnahR,nahR,NahR],
input =[SA~NahR],
output =[NahR]
){
PROCESS p1 = ConstitutiveProteinExpression(promoter
= PnahR, gene = nahR, protein = NahR)
}
|
CELLs and REGIONs¶
IBL allows for physical encapsulation of molecules, processes, and devices into CELL entities, as well as co-location of molecules, rules, processes, and cells into REGION entitis. By default, cells and regions physically separate molecules by introducing boundaries. However, IBL supports mechanisms to define translocation of molecules from cells into regions and vice versa, which allows for expressing secretion and uptake processes as well as cell-to-cell communication.
The example below illustrates the syntax used for:
- defining a cell, containing the AHL modulecule and a rule;
- defining a region, containing the AHL molecule and instantiating the cell defined above.
1 2 3 4 5 6 7 8 9 | DEFINE myCell TYPEOF CELL {
AHL = MOLECULE ()
RULE ahl_permeation: AHL <-> OUTSIDE
}
DEFINE well TYPEOF REGION {
AHL = MOLECULE (concentration=100 mM)
cell = myCell()
}
|
Verification Statements¶
IBL allows the designer to incorporate verification statements info various compartment types, e.g processes, devices, cells. The syntax of expressing the verification statements (also refered to as properties) is close to natural language and allow various patterns to be used.
Given below is a set of verification statements, illustrating the various patterns permitted by IBL:
1 2 3 4 5 | VERIFY property EVENTUALLY HOLDS // property will eventually become true, and then remains true
VERIFY property ALWAYS HOLDS // property is always true
VERIFY property NEVER HOLDS // property is never true
VERIFY property HOLDS IN STEADY-STATE // property will become true in steady-state
VERIFY EXPECTED [RFP] AT 1000 s IS ?
|
Biocompilation Directives¶
Biocompilation directives can also be incorporated into the various compartmetns of the IBL model, using the syntax below:
1 | ATGC ARRANGE : nahR, PnahR, Psal
|